Current Research
The Pickering Lab uses tools from epidemiology, genomics, and engineering to understand enteric disease transmission pathways among households in low-income settings and to develop effective, equitable, and scalable interventions to interrupt them.
Environmental Surveillance
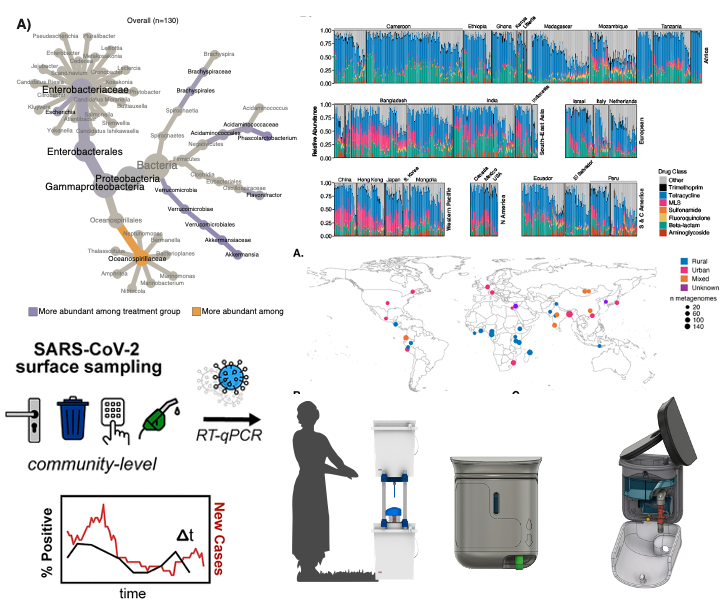
We use environmental sampling (surfaces, sewage, soil) to monitor trends in infectious disease in communities. Examples include soil surveillance for identifying communities with intestinal worm infections and high-touch surfaces for monitoring COVID-19 case trends.
Design, evaluate, and scale WASh Technologies
Leveraging human-centered design to co-create water, sanitation, and hygiene technologies with community partners is a focus in the lab. Examples include passive chlorination to increase access to safe drinking water and frugal foaming soap dispensers. We focus on technologies that avoid burdening target users.
infectious disease at the human-environment interface
We exploit long-read metagenomics and other molecular tools to track transmission of pathogens and antibiotic resistance (mobile elements, resistant bacterial strains) across humans, animals, and the environment.